Yeasts are a type of fungus that many of us associate with the production of alcoholic beverages and fermented foods. But there is a lot more to this complex group of microorganisms than just food and drink. Within this group, lurks yeasts that are harmful to human health: the so-called pathogens.
Our goal is to find and monitor these dangerous organisms in a precious natural resource: river water.
Refining river water quality monitoring for pathogenic yeast surveillance and exposure risk assessment: Science For Africa Foundation grant number: SFA-R16-102
South African Rivers
In South Africa, failing wastewater treatment plants and bad human practices are polluting our rivers – posing a health risk. The risk of exposure to these polluted waters, further increases due to natural disasters such as floods – which are made worse by climate change.
Hazards and Risks
Among the chemicals, bacteria, viruses, and parasites (pollutants in the river water) are the understudied and overlooked fungi - including the pathogenic yeasts. Yeasts frequently isolated from river water have represented members of Candida and other non-Candida species.
Vulnerable Communities
The proximity of communities situated near rivers increases their chances of exposure to the river water via general usage or flooding events. This makes them vulnerable to the pollutants found therein, which includes the understudied threat of pathogenic yeasts.
Meet the Scientists
Dr Cleo Conacher

Ms Heidi Steffen

Prof Alf Botha

What do we do?
As microbiologists we identify the small creatures present in the environment. River water is a particularly important medium for us as it might pose a health risk to those who come into contact with it. While others study microbes in various environments, we pay attention to the risks pertaining to pathogenic yeasts detected in river water.
We are qualified in the study of microorganisms and their interactions with the environment. We specifically focus on yeasts - with this project zooming into those that pose a health risk to humans. We also investigate yeast physiology and try to understand better how much damage they would be able to cause in humans.
Our research group actively started looking into the presence of pathogenic yeasts in river water back in 2020. We also investigated the development of a quantitative microbial risk assessment (QMRA) methodology. This is where we started to notice the specific limitations in the methodology used to detect and quantify pathogenic yeasts for the purpose of describing risk.
Apart from studying these yeasts and detecting them in the environment, we need a realistic understanding of community exposure scenarios. In our provisional risk assessment framework, we had to make assumptions for exposure paths. But, by engaging with communities in close proximity to the Eerste River, we are learning that there are exposure scenarios (previously overlooked) which more accurately describe risk.
Research focus
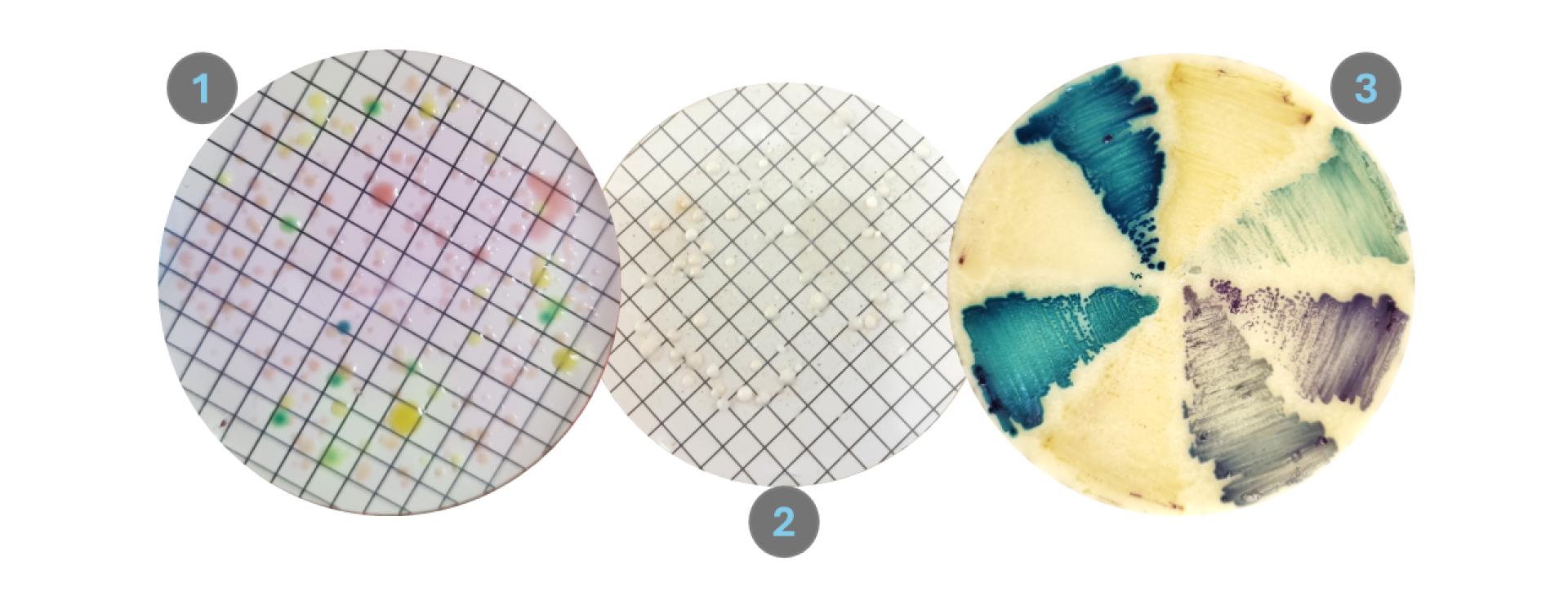
We perform culture-based methods to isolate yeasts from the environment. We do this by filtering river water through a gridded membrane (as seen on images 1 and 2) which capture the yeasts. These membranes are placed onto specified media (microbe food) to allow for microbial growth. Specific components in these media selectively allow for the growth of different microbes. Some components also initiate metabolic reactions that produce pigmented cell colonies.
While our interest is in the yeasts, we are still interested in the level of microbial pollution that can be quantified based on standard practices. The chromogenic medium Membrane Lactose Glucuronide Agar (MLGA) allows for identification of microbes associated with faecal matter: coliforms (yellow) and specifically Escherichia coli (green). Counting their colonies (dots of colour) provide an indication of how polluted the water is due to faecal contamination.
Isolating our yeasts occurs simultaneously with quantification of bacteria (on the left). However a different medium is used and we need to add components that prevent bacteria from growing: Sabauroud Dextrose Agar (SDA) with antibiotics. The cream and white dots are all yeast colonies that may represent various different species - but their colony morphology is not enough for us to identify them!
We isolate many yeasts from the river water. And to efficiently screen their identities we try to group them by the morphology they present on a selective chromogenic medium. Similar to MLGA, Candida Brilliance (CB) Agar induces colony colour differentiation. This helps us to group the yeasts based on morphology and then identify using more sensitive and definitive techniques.
We are hard at work developing better, more sensitive methods to detect and accurately quantify our target yeast species for the purpose of assessing health risks. This involves tackling the limitations we have identified during our previous research:
- Limited understanding of strain variability in virulence
- Accurate methods for quantifying yeast cells
- Links between environmental conditions and yeast prevalence/presence
- Realistic exposure scenarios that represent true contact with river water as experienced in communities
Pathogenic yeast species have reasons for being considered dangerous. What we aim to do is understanding what in their DNA might explain that danger. We are currently busy evaluating the different virulence phenotypes of Candida albicans and seeing if there are links to specific differences in their genetic make-up. So far, we have established that this yeast is quite diverse, and we believe that this diversity has been overlooked - meaning we do not have the right idea of how dangerous the specific strains are that occur in river water. We are developing molecular techniques that keep this in mind, such as qPCR (see link below).
This is a method that can be used for quantification of microorganisms. We are still in the development stages for this method as qPCR for quantification of yeasts have not been standardised. Along with this method we are trying to incorporate quantifying the live yeast cells only by using chemicals during the process to eliminate dead cells. This is called viability-qPCR. Promising preliminary results of this show that we have a candidate chemical compound to use; but we still need to evaluate whether this method is appropriate for our sampling protocol.
Together with understanding the microbial pollution levels of the river water, we are curious about the physico-chemical parameters of the water, which can tell us more about the interaction between these yeasts and their immediate environment. This is important because changes in these parameters could be a result of climate change or pollution events. The physical and chemical conditions also have an effect on the yeast's physiology - which might change how it interacts when it comes into contact with people. That is why we are busy developing an integrated biosensor in collaboration with colleagues in the Department of Electrical and Electronic Engineering, that would measure these conditions while simultaneously sampling water to perform the already described methods. It creates real-time snapshots of what the environment is like, and which microorganisms are present during that snapshot.
Thus far we have engaged with a community to get a better understanding of what types of exposures they experience with river water. These engagements, that we performed as workshops, were used to create a structured survey that would help us understand specific scenarios with additional information on frequency of exposure.
About our Funders and Collaborators
Science For Africa Foundation
This project is funded by the Science For Africa (SFA) Foundation, under the Grand Challenges Africa scheme, in the focus area of: Accelerating Catalyzing Solutions for Climate Change's Impact on Health and Gender. SFA Foundation implements funding for Africa-led research and innovation that has potential for impact across the globe. Their support has been critical to launch this research.
Water Research Commission
This project is co-funded by the Water Research Commission of South Africa within the thematic area of Water Quality and Health, under grant number 2025/2026-01818. The Water Research Commission is a governmental organisation that supports research and innovation in the local water sector. We are proud to continue the work of this project with their support.
Collaborative Governance (Co-Go) for Water Security
Co-Go has been instrumental in building the bridges for communication between our research group and local communities. The connections between community, industry, and government, predominantly achieved by Dr Charon Buchner-Marais’ immense networking, has allowed our research to move beyond the bounds of a laboratory and into real-world systems.
Electrical and Electronic Engineering (SU)
On the opposite side of Banghoek road, Dr Taskeen Ebrahim and her team in Electrical and Electronic Engineering are hard at work advancing the technical aspects of the biosensor-sampler prototype we envision using in the field. Their engineering prowess and passion for impact-led research has allowed our prototypes to advance by leaps and bounds.